

Nucleotide excision repair (NER)
|
|
| The nucleotide excision repair
(NER) pathway is responsible for removing a wide range of damage from DNA,
including UV-induced CPDs and 6-4PPs. The ability of this mechanism
to recognise so many different forms of DNA damage arises from its targeting
of distortions in the DNA double helix instead of (or in addition to) identifying
the specific chemistry of individual lesions [1]. The basic process
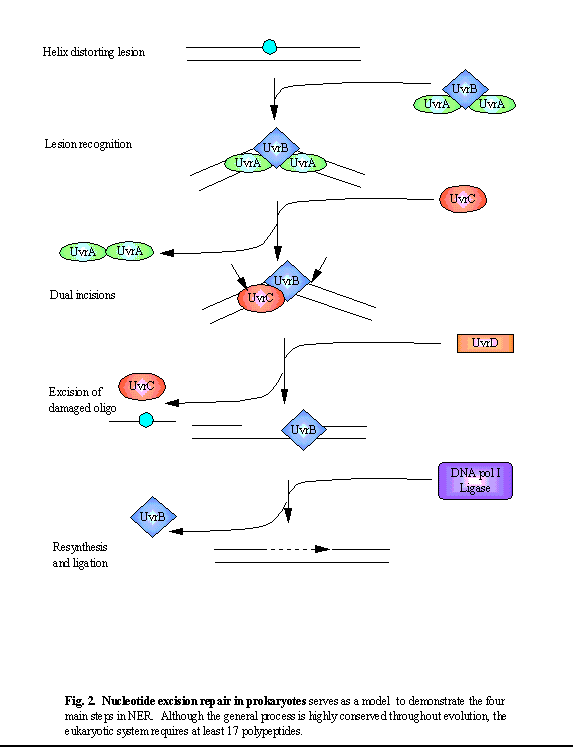
of NER consists of four steps (see figure below):
lesion recognition, incisions on the damaged strand on either side of the
lesion, excision of the damaged oligonucleotide, and resynthesis (and ligation)
of the excised fragment, using the undamaged strand as a template.
Although NER protein homology
is strongly conserved within eukaryotes, there is very little homology
between the prokaryotic NER proteins and their eukaryotic counterparts.
However, the general process of NER is conserved, and a review of the prokaryotic
model is useful in understanding the (more complicated and less well defined)
NER processes in eukaryotes.
|
| Although NER is the major mechanism for repair of UV-induced photoproducts in DNA, alternative mechanisms exist in some species which can also repair these lesions. |
|
|
| Prokaryotic NER |
| NER in E. coli, requires the
products of only six genes, of which three, UvrA, UvrB and UvrC, comprise
the so-called (A)BC excinuclease complex, although a complex between all
three does not exist at the same time (see figure).
1. Initial lesion recognition is performed by a complex consisting of two UvrA and one UvrB subunits. This A2B1 complex tracks along the DNA backbone, recognising a lesion first through the UvrA dimer which is specific for damage recognition on double-stranded DNA [2]. Binding of the complex to the lesion activates the UvrB-dependent helicase activity, leading to an unwinding/kinking of the DNA. This allows further recognition of the damaged strand by UvrB, which is specific for damaged single-stranded DNA [3]. Such a multistep damage recognition process allows for both specificity and a wide substrate range. The conformational change induced by the tight binding of UvrB weakens its association with the UvrA dimer. Dissociation of the complex leaves a stable UvrB-DNA complex [4, 5]. 2. The binding of UvrC to the UvrB-DNA complex initiates incisions on the damaged DNA strand: first, at the 4th or 5th phosphodiester bond 3' to the lesion (mediated by UvrB) and a few seconds later at the 8th phosphodiester bond 5' (mediated by UvrC), leading to the formation of a 12-13 nt long oligomer [6]. 3. The subsequent binding of UvrD (helicase II) dissociates the oligomer from the duplex DNA, making the site accessible to other enzymes. 4. The repair specific DNA
polymerase I (pol I) synthesises the repair patch, and DNA ligase then
completes the repair process.
|
|
| Transcription-coupled repair (TCR) in prokaryotes |
| In addition to this generalised
repair system which acts to remove damage from throughout the entire genome,
an additional NER sub-pathway exists which allows for the preferential
repair of (transcription-blocking) lesions on the transcribed strand of
active genes [7-11]. In E. coli, this transcription-coupled
repair pathway is mediated by the product of the mfd gene, which
has been termed the transcription repair coupling factor (TRCF).
This coupling factor displaces RNA pol stalled at lesions and recruits
UvrA to the damage site, facilitating repair of the lesion [12].
|
|
Examination of cells from patients with genetic disorders compromising their NER capability have been invaluable in understanding NER processes in human cells. |
|
|

|
|
| References |
| 1. Gunz, D., M.T. Hess, and H. Naegeli, Recognition of DNA adducts
by human nucleotide excision repair. Evidence for a thermodynamic probing
mechanism. Journal of Biological Chemistry, 1996. 271(41): p. 25089-98.
2. Seeberg, E. and R.P. Fuchs, Acetylaminofluorene bound to different guanines of the sequence -GGCGCC- is excised with different efficiencies by the UvrABC excision nuclease in a pattern not correlated to the potency of mutation induction. Proc Natl Acad Sci U S A, 1990. 87(1): p. 191-4. 3. Hsu, D.S., et al., Structure and function of the UvrB protein. Journal of Biological Chemistry, 1995. 270(14): p. 8319-27. 4. Visse, R., et al., Analysis of UvrABC endonuclease reaction intermediates on cisplatin-damaged DNA using mobility shift gel electrophoresis. Journal of Biological Chemistry, 1992. 267(10): p. 6736-42. 5. Delagoutte, E., et al., Sequence-dependent modulation of nucleotide excision repair: the efficiency of the incision reaction is inversely correlated with the stability of the pre-incision UvrB-DNA complex. Journal of Molecular Biology, 1997. 266(4): p. 703-10. 6. Lin, J.J. and A. Sancar, Reconstitution of nucleotide excision nuclease with UvrA and UvrB proteins from Escherichia coli and UvrC protein from Bacillus subtilis. J Biol Chem, 1990. 265(34): p. 21337-41. 7. Mellon, I. and P.C. Hanawalt, Induction of the Escherichia coli lactose operon selectively increases repair of its transcribed DNA strand. Nature, 1989. 342(6245): p. 95-8. 8. Selby, C.P. and A. Sancar, Molecular mechanism of transcription-repair coupling. Science, 1993. 260(5104): p. 53-8. 9. Selby, C.P. and A. Sancar, Gene- and strand-specific repair in vitro: partial purification of a transcription-repair coupling factor. Proceedings of the National Academy of Sciences of the United States of America, 1991. 88(18): p. 8232-6. 10. Selby, C.P., E.M. Witkin, and A. Sancar, Escherichia coli mfd mutant deficient in "mutation frequency decline" lacks strand-specific repair: in vitro complementation with purified coupling factor. Proceedings of the National Academy of Sciences of the United States of America, 1991. 88(24): p. 11574-8. 11. Selby, C.P. and A. Sancar, Transcription-repair coupling and mutation frequency decline. Journal of Bacteriology, 1993. 175(23): p. 7509-14. 12. Selby, C.P. and A. Sancar, Structure and function of transcription-repair
coupling factor. II. Catalytic properties. Journal of Biological Chemistry,
1995. 270(9): p. 4890-5.
|
|
|
